Quantitative Imaging At Nanoscale by SUper-resolution (QIAN SU LAB)
- to bridge biomedical engineering and biophysical nanotools with fundamental sciences, by applying advanced imaging methods to quantitatively pinpoint the molecular secrets during important biological processes
Multi-functional Imaging Toolbox for Intracellular Dynamics
To be updated soon... ...
Novel in vitro reconstitution assay for Autophagy and Mitochondrial Metabolism
|
I have longstanding interest in understanding autolysosome composition and dynamics, the fundamental autophagy (literally, “self-eating” in Greek) mechanism implicated in cancer, neurodegenerative diseases, infectious diseases, and metabolic diseases and awarded Nobel Prize 2017.
From 2011 to 2016, by leading 3 PhDs and 2 RAs from Peking, Tsinghua University and Chinese Academy of Science, I have established an easy-to-use single-molecule in vitro reconstitution assay to study motor protein induced membrane deformation, reformation and dynamics. This in vitro pure system provides an unprecedented flexibility in manipulating motor protein conformation, lipid composition, vesicle size, Ca2+ion concentration and ATP/ADP binding condition, which cannot be genetically or physiologically controlled in intracellular environment. Notably, Development Cell Editor Dr Ilil Carmi commented our work as “an extremely convenient in vitro knock-down treatment could be achieved through the reconstitution assay” [1]. Using this powerful in vitro experimental platform, our team have demonstrated that the kinesin-1 KIF5B, through interactions with PtdIns(4,5)P2, drives tubulation by pulling on the autolysosome membrane, revealing a motor-based membrane deformation process that helps maintain lysosomal homeostasis [1,2]. This work was recommended and highlighted by F1000 Prime by Prof. Wei GUO and Prof. Bin WU: “This paper provides a working model to explain how microtubule motor proteins are involved in lysosome membrane dynamics.” A following work using this in vitro reconstitution system for mitochondrial metabolism, uncovered an essential mechanism for mitochondrial network formation driven by KIF5B and mediated by MFN1/2 and revealed that different parts of the mitochondrial network are formed by different mechanisms [3], cover story in Cell Research. The well-established in vitro reconstitution assay with single-molecule sensitivity has further been applied to understand CapZ and actin in autophagy [4], ATP/ADP in kinesin motility [5], Ca2+ in myosin motion [6], Golgi derived vesicles [7] and I was listed as contributing authors in these papers. Three more invited book chapter and review papers have been published in the following years [8-10]. |
Pinpoint Chromatin 3D Structure and DNA Replication by Super-resolution Microscopy
|
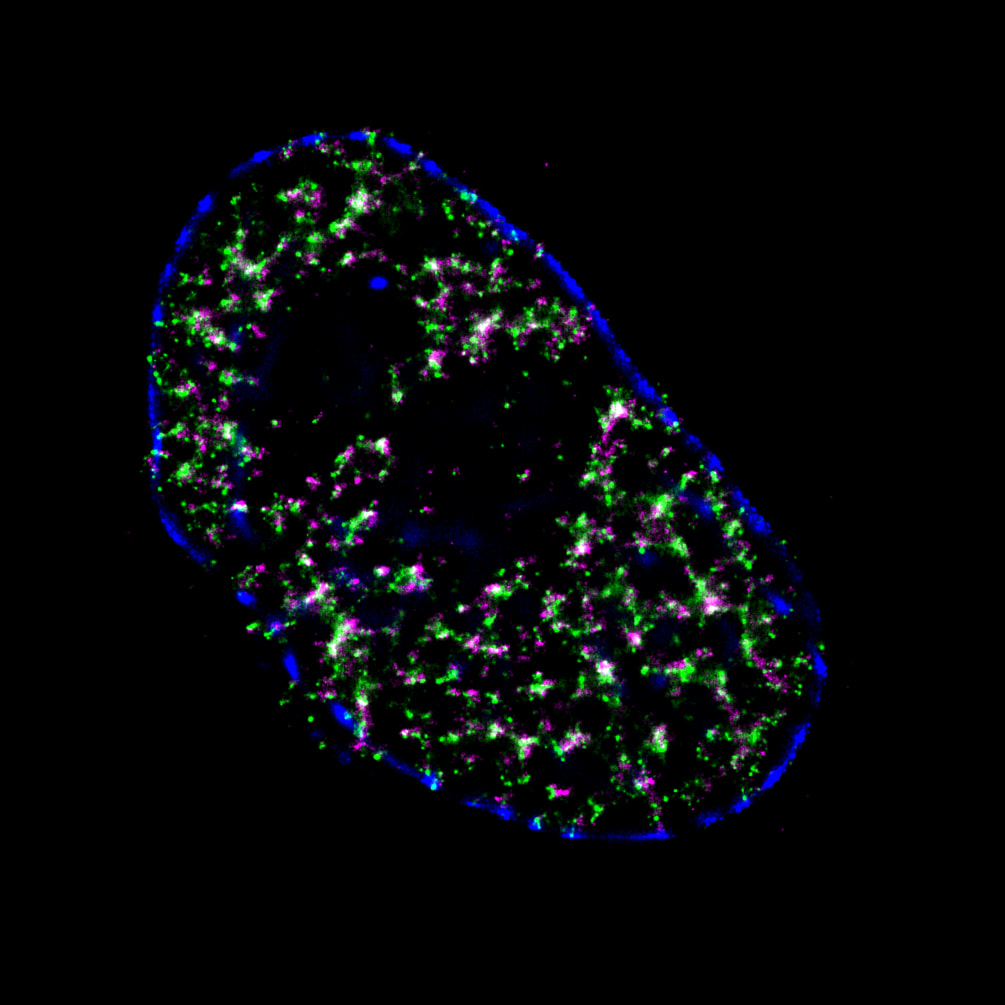
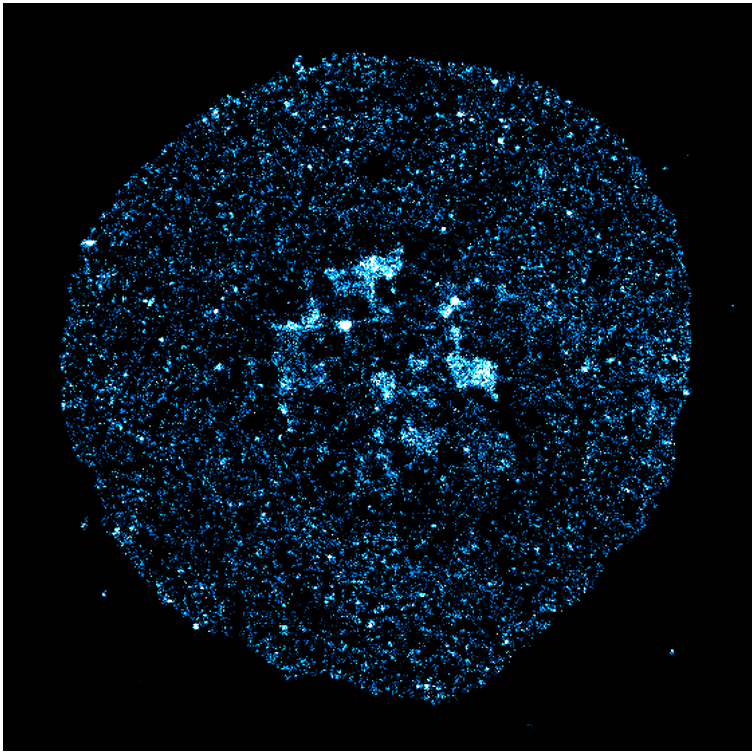
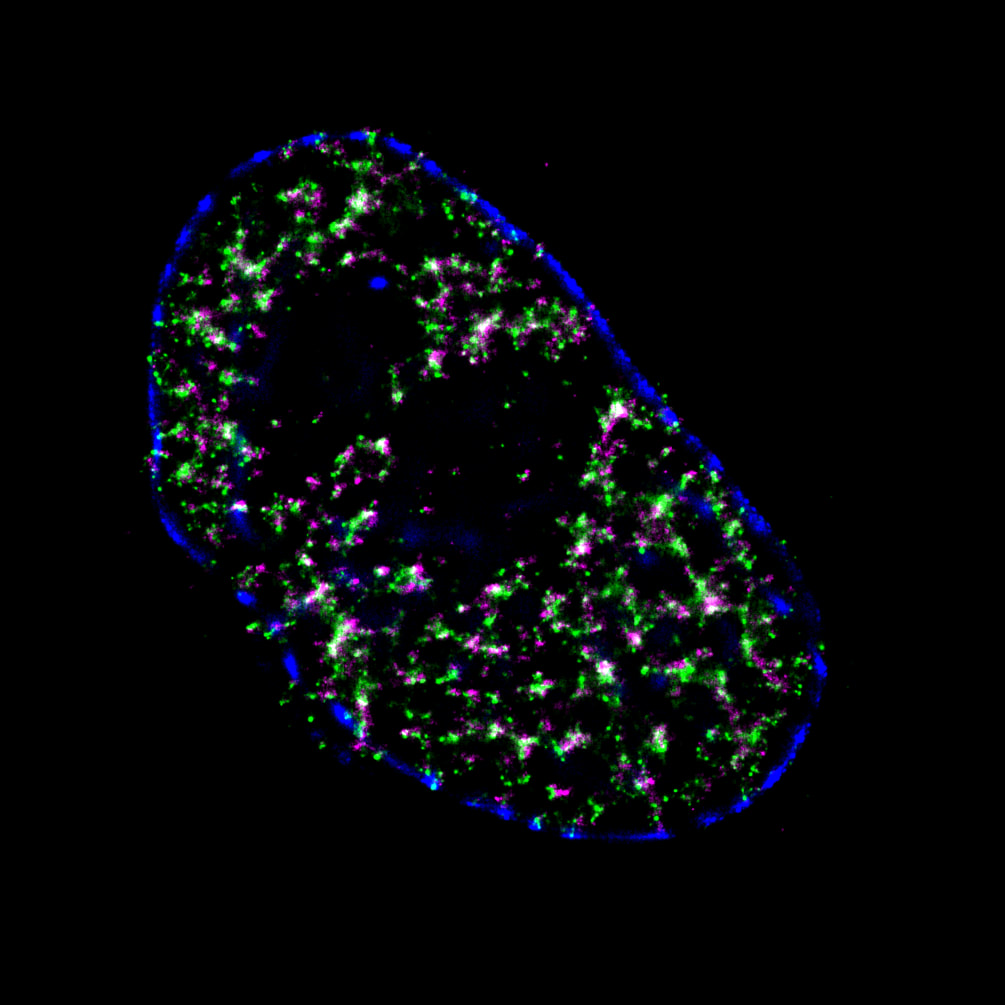
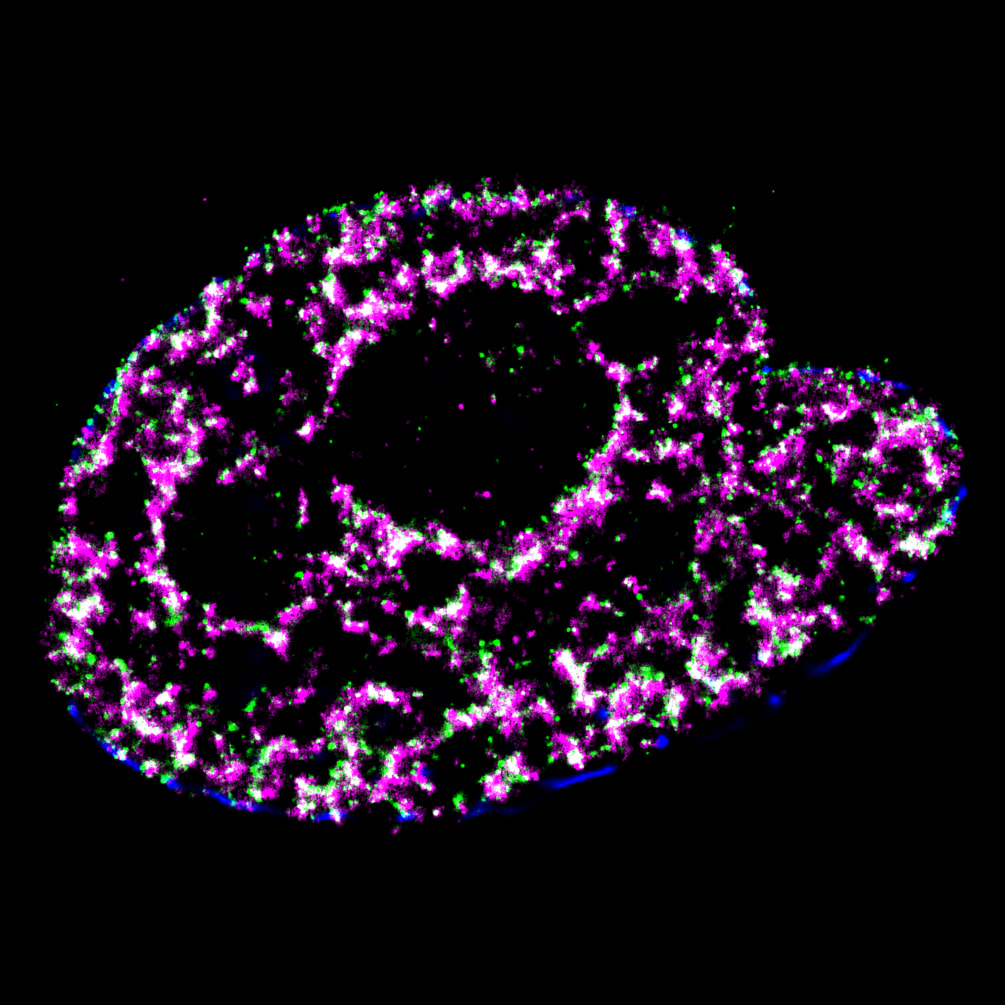
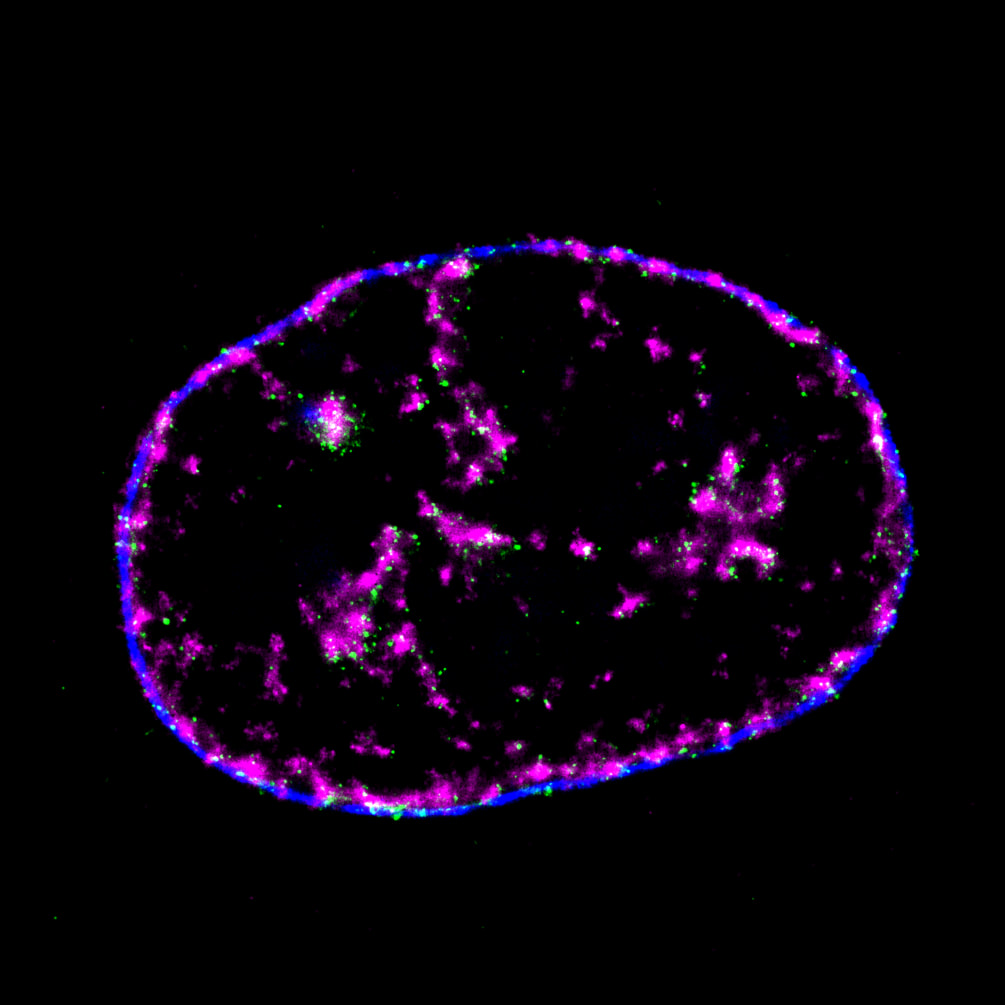
Mammalian DNA replication is initiated at numerous replication origins, which are clustered into thousands of replication domains (RDs) across the genome. However, it remains unclear whether the replication origins within each RD are activated stochastically. To understand how replication is regulated at the sub-RD level, I have worked with Prof Xiaoliang Sunney XIE and Prof Xiaowei ZHUANG and their colleagues at Harvard and developed multi-colour 3D STORM chromatin imaging analysis. This new method advanced the field by allowing one to visualise the spatio-temporal organization, morphology, and in situ epigenetic signatures of individual replication foci (RFi) across S-phase. Importantly, I have revealed a hierarchical radial pattern of RFi propagation that reverses its directionality from early to late S-phase, and is diminished upon caffeine treatment or CTCF knockdown. Together with simulation and bioinformatics analyses, my research works point to a ‘CTCF-organized REplication Propagation’ (CoREP) model. The CoREP model suggests a non-random selection mechanism for replication activation mediated by CTCF at the sub-RD level, as well as the critical involvement of local chromatin environment in regulating replication in space and time [11].
The well-established super-resolution assay (optical configuration and analysis algorithms) with 30nm resolution in cell nucleus has also been applied to understand the role of PTEN protein in DNA repair [12], DNA oligo bio-conjugation and localization [13] and transcription factor in bacterial cells [14]. |
Visualise integrin αIIbβ3 dynamics and its interactome in platelet clotting
|
By working with Dr JU and Prof JACKSON from University of Sydney, I have established the TIRF/HiLoM/ STORM system at Charles Perkins Centre of USYD/HRI. With a palette of new organic probes and dye-conjugated antibodies, we have mapped the spatio-temporal distribution of integrin αIIbβ3 with “bent” vs. “extended” conformation attributed to “resting” vs. “activated” platelets respectively at single cell level. Furthermore, I performed Multi-colour SM imaging to correlate these integrin conformation states with dynamics of cell membrane, actin, and microtubules. This interactome analysis demonstrates that the resting integrin distribute ahead of membrane spreading followed by the activated integrin in sequence, suggesting the role of integrin activation in leading the molecular machinery during the platelet spreading and aggregation process.
My further STORM analysis further mapped integrin αIIbβ3 conformational landscape across different stages of platelet activation at single-molecule resolution (~20 nm). The molecular “fingerprints” obtained by my SM/SR imaging approaches will conceptually advance the knowledge on integrin structure, affinity regulation and dynamics which sheds light to novel integrin targeting therapeutics in future. |